-
Notifications
You must be signed in to change notification settings - Fork 0
/
qfeatures_intro.html
380 lines (290 loc) · 13.9 KB
/
qfeatures_intro.html
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23
24
25
26
27
28
29
30
31
32
33
34
35
36
37
38
39
40
41
42
43
44
45
46
47
48
49
50
51
52
53
54
55
56
57
58
59
60
61
62
63
64
65
66
67
68
69
70
71
72
73
74
75
76
77
78
79
80
81
82
83
84
85
86
87
88
89
90
91
92
93
94
95
96
97
98
99
100
101
102
103
104
105
106
107
108
109
110
111
112
113
114
115
116
117
118
119
120
121
122
123
124
125
126
127
128
129
130
131
132
133
134
135
136
137
138
139
140
141
142
143
144
145
146
147
148
149
150
151
152
153
154
155
156
157
158
159
160
161
162
163
164
165
166
167
168
169
170
171
172
173
174
175
176
177
178
179
180
181
182
183
184
185
186
187
188
189
190
191
192
193
194
195
196
197
198
199
200
201
202
203
204
205
206
207
208
209
210
211
212
213
214
215
216
217
218
219
220
221
222
223
224
225
226
227
228
229
230
231
232
233
234
235
236
237
238
239
240
241
242
243
244
245
246
247
248
249
250
251
252
253
254
255
256
257
258
259
260
261
262
263
264
265
266
267
268
269
270
271
272
273
274
275
276
277
278
279
280
281
282
283
284
285
286
287
288
289
290
291
292
293
294
295
296
297
298
299
300
301
302
303
304
305
306
307
308
309
310
311
312
313
314
315
316
317
318
319
320
321
322
323
324
325
326
327
328
329
330
331
332
333
334
335
336
337
338
339
340
341
342
343
344
345
346
347
348
349
350
351
352
353
354
355
356
357
358
359
360
361
362
363
364
365
366
367
368
369
370
371
372
373
374
375
376
377
378
379
380
<!DOCTYPE html>
<html lang="" xml:lang="">
<head>
<title>Handling quantitative proteomics features?</title>
<meta charset="utf-8" />
<meta name="author" content="Laurent Gatto and Christophe Vanderaa" />
<script src="qfeatures_intro_files/header-attrs-2.10/header-attrs.js"></script>
<script src="qfeatures_intro_files/xaringanExtra-webcam-0.0.1/webcam.js"></script>
<script id="xaringanExtra-webcam-options" type="application/json">{"width":"200","height":"200","margin":"1em"}</script>
<link href="qfeatures_intro_files/tile-view-0.2.6/tile-view.css" rel="stylesheet" />
<script src="qfeatures_intro_files/tile-view-0.2.6/tile-view.js"></script>
<script src="qfeatures_intro_files/xaringanExtra_fit-screen-0.2.6/fit-screen.js"></script>
<link href="qfeatures_intro_files/xaringanExtra-extra-styles-0.2.6/xaringanExtra-extra-styles.css" rel="stylesheet" />
<link href="qfeatures_intro_files/panelset-0.2.6/panelset.css" rel="stylesheet" />
<script src="qfeatures_intro_files/panelset-0.2.6/panelset.js"></script>
<script src="qfeatures_intro_files/htmlwidgets-1.5.3/htmlwidgets.js"></script>
<script src="qfeatures_intro_files/jquery-3.5.1/jquery.min.js"></script>
<link href="qfeatures_intro_files/datatables-css-0.0.0/datatables-crosstalk.css" rel="stylesheet" />
<script src="qfeatures_intro_files/datatables-binding-0.18/datatables.js"></script>
<link href="qfeatures_intro_files/dt-core-1.10.20/css/jquery.dataTables.min.css" rel="stylesheet" />
<link href="qfeatures_intro_files/dt-core-1.10.20/css/jquery.dataTables.extra.css" rel="stylesheet" />
<script src="qfeatures_intro_files/dt-core-1.10.20/js/jquery.dataTables.min.js"></script>
<link href="qfeatures_intro_files/crosstalk-1.1.1/css/crosstalk.css" rel="stylesheet" />
<script src="qfeatures_intro_files/crosstalk-1.1.1/js/crosstalk.min.js"></script>
<link rel="stylesheet" href="xaringan-themer.css" type="text/css" />
</head>
<body>
<textarea id="source">
class: center, middle, inverse, title-slide
# Handling quantitative proteomics features?
### Laurent Gatto and Christophe Vanderaa
### 2021/08/10
---
class: middle
name: cc-by
### Get the slides at http://bit.ly/202108SCP
These slides are available under a **creative common
[CC-BY license](http://creativecommons.org/licenses/by/4.0/)**. You are
free to share (copy and redistribute the material in any medium or
format) and adapt (remix, transform, and build upon the material) for
any purpose, even commercially
<img height="20px" alt="CC-BY" src="https://raw.githubusercontent.com/UCLouvain-CBIO/scp-teaching/main/img/cc1.jpg" />.
---
class: middle, center, inverse
# What is QFeatures?
---
class: middle
### What is QFeatures?
- An R/[Bioconductor](http://www.bioconductor.org/packages/QFeatures)
*package* to process and manage quantitative proteomics data.
- A *class*, i.e. the container that will hold the quantitative
(single cell) proteomics data and that will **track** and **record**
the processing steps.
---
class: middle, center, inverse
# Feature aggregation in proteomics
---
class: middle
.pull-left[
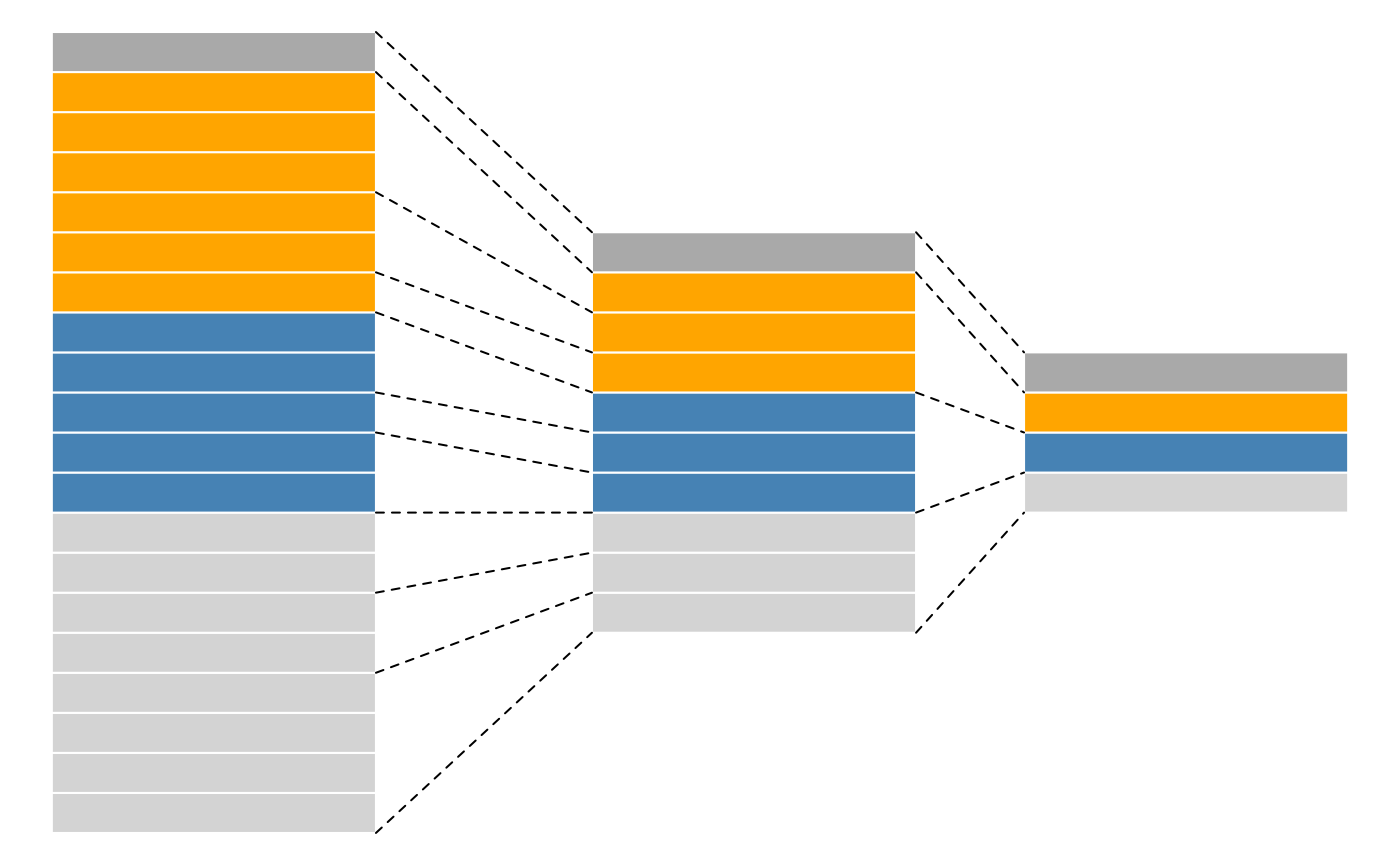
### Feature aggregation in proteomics
- One or more PSMs are aggregated into peptides.
- One or more peptides are aggregated into protein groups.
**QFeatures** does this in a consistent and reproducible way and
records these steps, so that one can easily **track** features across
levels.
]
.pull-right[
<!-- -->
]
---
class: middle, center, inverse
# What happens under the hood?
---
class:
.panelset[
.panel[.panel-name[QFeatures]
A **QFeatures** object containing three **assays**, named PSM1, PSM2
and PSM3. Each assay contains quantitative data for a set of features
(rows) and samples (columns). The **QFeatures** object also contains
sample annotations matching columns across all assays.
<img src="./figs/qfeatures_intro_1.svg" width="70%" style="display: block; margin: auto;" />
Such a **QFeatures** object can be created with the [`readSCP()`]()
function.
]
.panel[.panel-name[assay]
<div id="htmlwidget-e05e34fd5962cf6a2c69" style="width:100%;height:auto;" class="datatables html-widget"></div>
<script type="application/json" data-for="htmlwidget-e05e34fd5962cf6a2c69">{"x":{"filter":"none","data":[["PSM1","PSM2","PSM3","PSM4","PSM5","PSM6","PSM7","PSM8","PSM9","PSM10"],[1,2,3,4,5,6,7,8,9,10],[11,12,13,14,15,16,17,18,19,20],["","","","","","","","","",""],["SYGFNAAR","SYGFNAAR","SYGFNAAR","ELGNDAYK","ELGNDAYK","ELGNDAYK","IAEESNFPFIK","IAEESNFPFIK","IAEESNFPFIK","IAEESNFPFIK"],["ProtA","ProtA","ProtA","ProtA","ProtA","ProtA","ProtB","ProtB","ProtB","ProtB"],[1,2,3,4,5,6,7,8,9,10],["Mitochondrion","Mitochondrion","Mitochondrion","Mitochondrion","Mitochondrion","Mitochondrion","unknown","unknown","unknown","unknown"],[0.084,0.077,0.063,0.073,0.012,0.011,0.075,0.038,0.028,0.097]],"container":"<table class=\"display\">\n <thead>\n <tr>\n <th> <\/th>\n <th>S1<\/th>\n <th>S2<\/th>\n <th>...<\/th>\n <th>Sequence<\/th>\n <th>Protein<\/th>\n <th>Var<\/th>\n <th>location<\/th>\n <th>pval<\/th>\n <\/tr>\n <\/thead>\n<\/table>","options":{"columnDefs":[{"className":"dt-right","targets":[1,2,6,8]},{"orderable":false,"targets":0}],"order":[],"autoWidth":false,"orderClasses":false}},"evals":[],"jsHooks":[]}</script>
]
.panel[.panel-name[colData]
<div id="htmlwidget-6b6e5dc69ade1cc055c3" style="width:100%;height:auto;" class="datatables html-widget"></div>
<script type="application/json" data-for="htmlwidget-6b6e5dc69ade1cc055c3">{"x":{"filter":"none","data":[["1","2","3","4","5","6","7","8","9","10","11","12","13","14","15","16","17","18","19","20","21","22","23","24","25","26","27","28","29","30","31","32","33","34","35","36"],["S1","S2","S3","S4","S5","S6","S7","S8","S9","S10","S11","S12","S13","S14","S15","S16","S17","S18","S19","S20","S21","S22","S23","S24","S25","S26","S27","S28","S29","S30","S31","S32","S33","S34","S35","S36"],["LC1","LC1","LC1","LC1","LC1","LC1","LC1","LC1","LC1","LC1","LC1","LC1","LC2","LC2","LC2","LC2","LC2","LC2","LC2","LC2","LC2","LC2","LC2","LC2","LC3","LC3","LC3","LC3","LC3","LC3","LC3","LC3","LC3","LC3","LC3","LC3"],["carrier","empty","cellA","cellB","cellC","blank","cellA","cellB","cellC","cellA","cellB","cellC","carrier","empty","cellA","cellB","cellC","blank","cellA","cellB","cellC","cellA","cellB","cellC","carrier","empty","cellA","cellB","cellC","blank","cellA","cellB","cellC","cellA","cellB","cellC"]],"container":"<table class=\"display\">\n <thead>\n <tr>\n <th> <\/th>\n <th>sample<\/th>\n <th>LC_batch<\/th>\n <th>type<\/th>\n <\/tr>\n <\/thead>\n<\/table>","options":{"order":[],"autoWidth":false,"orderClasses":false,"columnDefs":[{"orderable":false,"targets":0}]}},"evals":[],"jsHooks":[]}</script>
]
]
---
class:
#### Aggregating PSMs to peptides
<img src="./figs/qfeatures_intro_2.svg" style="display: block; margin: auto;" />
```r
qf <- aggregateFeaturesOverAssays(qf,
i = c("PSM1", "PSM2", "PSM3"), ## assays to be aggregated
name = c("Pep1", "Pep2", "Pep3"), ## new assay name
fcol = "peptide", ## what to aggregate
fun = colMedians) ## how to aggregate
```
---
class:
#### Joining peptide assays
<img src="./figs/qfeatures_intro_3.svg" style="display: block; margin: auto;" />
```r
qf <- joinAssays(qf,
c("Pep1", "Pep2", "Pep3"), ## assays to be joined
name = "Peptides") ## new assay name
```
---
class:
#### Aggregate peptides into proteins
<img src="./figs/qfeatures_intro_4.svg" style="display: block; margin: auto;" />
```r
qf <- aggregateFeatures(qf,
"Peptides", ## assay to be aggregated
"Protein", ## new assay name
name = "proteins", ## what to aggregate
fun = colMedians) ## how to aggregate
```
---
class: middle, center, inverse
# Why tracking assays
---
class: middle
<img src="https://rformassspectrometry.github.io/QFeatures/articles/QFeatures_files/figure-html/plotstat-1.png" width="90%" style="display: block; margin: auto;" />
---
class: middle, center, inverse
# More with QFeatures and scp
---
class: middle
### More with QFeatures and scp
- quality control
- feature and sample filtering, for example `filterFeatures()`
- transformation and normalisation, for example `normalize()`, `logTransform()`
- imputation with `impute()`
- batch correction
- visualisaton
- the full R/Bioconductor statistical learning toolkit
---
class: middle, center, inverse
# Exercise
---
class: middle
#### Match the code to the `QFeatures` object structure.
A.
<img src="./figs/qf_plot_qst1.png" width="30%" />
B.
<img src="./figs/qf_plot_qst2.png" width="30%" />
C.
<img src="./figs/qf_plot_qst3.png" width="30%" />
```r
1. joinAssays(); aggregateFeatures(); logTranform()
2. aggregateFeaturesOverAssays(); logTransform(); joinAssays()
3. logTransfrom(); joinAssays(); aggregateFeatures()
```
Connect to **http://www.wooclap.com/YYRQRM**
---
class: middle
### Further information
- The **QFeatures** webpage: http://RforMassSpectrometry.github.io/QFeatures
- The **scp** webpage: http://UCLouvain-CBIO.github.io/scp
- Online **tutotial**: https://lgatto.github.io/QFeaturesScpWorkshop2021/
### Funding
Fonds de la Recherche Scientifique (FNRS), Belgium
</textarea>
<style data-target="print-only">@media screen {.remark-slide-container{display:block;}.remark-slide-scaler{box-shadow:none;}}</style>
<script src="https://remarkjs.com/downloads/remark-latest.min.js"></script>
<script>var slideshow = remark.create({
"highlightStyle": "github",
"highlightLines": true,
"ratio": "16:9",
"countIncrementalSlides": true
});
if (window.HTMLWidgets) slideshow.on('afterShowSlide', function (slide) {
window.dispatchEvent(new Event('resize'));
});
(function(d) {
var s = d.createElement("style"), r = d.querySelector(".remark-slide-scaler");
if (!r) return;
s.type = "text/css"; s.innerHTML = "@page {size: " + r.style.width + " " + r.style.height +"; }";
d.head.appendChild(s);
})(document);
(function(d) {
var el = d.getElementsByClassName("remark-slides-area");
if (!el) return;
var slide, slides = slideshow.getSlides(), els = el[0].children;
for (var i = 1; i < slides.length; i++) {
slide = slides[i];
if (slide.properties.continued === "true" || slide.properties.count === "false") {
els[i - 1].className += ' has-continuation';
}
}
var s = d.createElement("style");
s.type = "text/css"; s.innerHTML = "@media print { .has-continuation { display: none; } }";
d.head.appendChild(s);
})(document);
// delete the temporary CSS (for displaying all slides initially) when the user
// starts to view slides
(function() {
var deleted = false;
slideshow.on('beforeShowSlide', function(slide) {
if (deleted) return;
var sheets = document.styleSheets, node;
for (var i = 0; i < sheets.length; i++) {
node = sheets[i].ownerNode;
if (node.dataset["target"] !== "print-only") continue;
node.parentNode.removeChild(node);
}
deleted = true;
});
})();
(function() {
"use strict"
// Replace <script> tags in slides area to make them executable
var scripts = document.querySelectorAll(
'.remark-slides-area .remark-slide-container script'
);
if (!scripts.length) return;
for (var i = 0; i < scripts.length; i++) {
var s = document.createElement('script');
var code = document.createTextNode(scripts[i].textContent);
s.appendChild(code);
var scriptAttrs = scripts[i].attributes;
for (var j = 0; j < scriptAttrs.length; j++) {
s.setAttribute(scriptAttrs[j].name, scriptAttrs[j].value);
}
scripts[i].parentElement.replaceChild(s, scripts[i]);
}
})();
(function() {
var links = document.getElementsByTagName('a');
for (var i = 0; i < links.length; i++) {
if (/^(https?:)?\/\//.test(links[i].getAttribute('href'))) {
links[i].target = '_blank';
}
}
})();
// adds .remark-code-has-line-highlighted class to <pre> parent elements
// of code chunks containing highlighted lines with class .remark-code-line-highlighted
(function(d) {
const hlines = d.querySelectorAll('.remark-code-line-highlighted');
const preParents = [];
const findPreParent = function(line, p = 0) {
if (p > 1) return null; // traverse up no further than grandparent
const el = line.parentElement;
return el.tagName === "PRE" ? el : findPreParent(el, ++p);
};
for (let line of hlines) {
let pre = findPreParent(line);
if (pre && !preParents.includes(pre)) preParents.push(pre);
}
preParents.forEach(p => p.classList.add("remark-code-has-line-highlighted"));
})(document);</script>
<script>
slideshow._releaseMath = function(el) {
var i, text, code, codes = el.getElementsByTagName('code');
for (i = 0; i < codes.length;) {
code = codes[i];
if (code.parentNode.tagName !== 'PRE' && code.childElementCount === 0) {
text = code.textContent;
if (/^\\\((.|\s)+\\\)$/.test(text) || /^\\\[(.|\s)+\\\]$/.test(text) ||
/^\$\$(.|\s)+\$\$$/.test(text) ||
/^\\begin\{([^}]+)\}(.|\s)+\\end\{[^}]+\}$/.test(text)) {
code.outerHTML = code.innerHTML; // remove <code></code>
continue;
}
}
i++;
}
};
slideshow._releaseMath(document);
</script>
<!-- dynamically load mathjax for compatibility with self-contained -->
<script>
(function () {
var script = document.createElement('script');
script.type = 'text/javascript';
script.src = 'https://mathjax.rstudio.com/latest/MathJax.js?config=TeX-MML-AM_CHTML';
if (location.protocol !== 'file:' && /^https?:/.test(script.src))
script.src = script.src.replace(/^https?:/, '');
document.getElementsByTagName('head')[0].appendChild(script);
})();
</script>
</body>
</html>